招贤纳士
研究生招生
欢迎具有生物信息学、生物化学和分子生物学、基础医学等相关背景且热爱科学研究的同学报考硕士和博士研究生。有意者请发详细个人简历到本人邮箱提前联系。
我们将提供最好的科研实验条件,瞄准国际前沿的研究课题,系统全面训练研究生的综合科研能力。鼓励研究生参加国内外的学术会议以提高专业素质,以使每一位研究生有机会作出第一流的研究工作,努力为你们未来职业发展提供最好的平台。
研究生招生信息详见中山大学研究生招生网
招聘博士后和专职科研人员
申请者基本条件
- 有相关方向的博士学位
- 对科学研究具有强烈的兴趣,工作认真负责,有优良的科学道德
- 积极乐观,有良好的团队协作精神
- 有较强的中英文阅读、写作能力
专业背景要求(符合任何一项均可申请)
- 有生物信息学、基因组学相关背景并掌握熟练的相关编程技能(如R,python,perl等),有大规模二代测序数据分析经验的优先;
- 具备分子生物学,细胞生物学或者生物化学背景,并备具较强的相关实验生物学技能。
基本职责
- 独立开展相关课题研究
- 协助课题组长指导研究生共同开展研究活动
- 协助课题组长进行项目申请以及后续项目管理
岗位及相关待遇
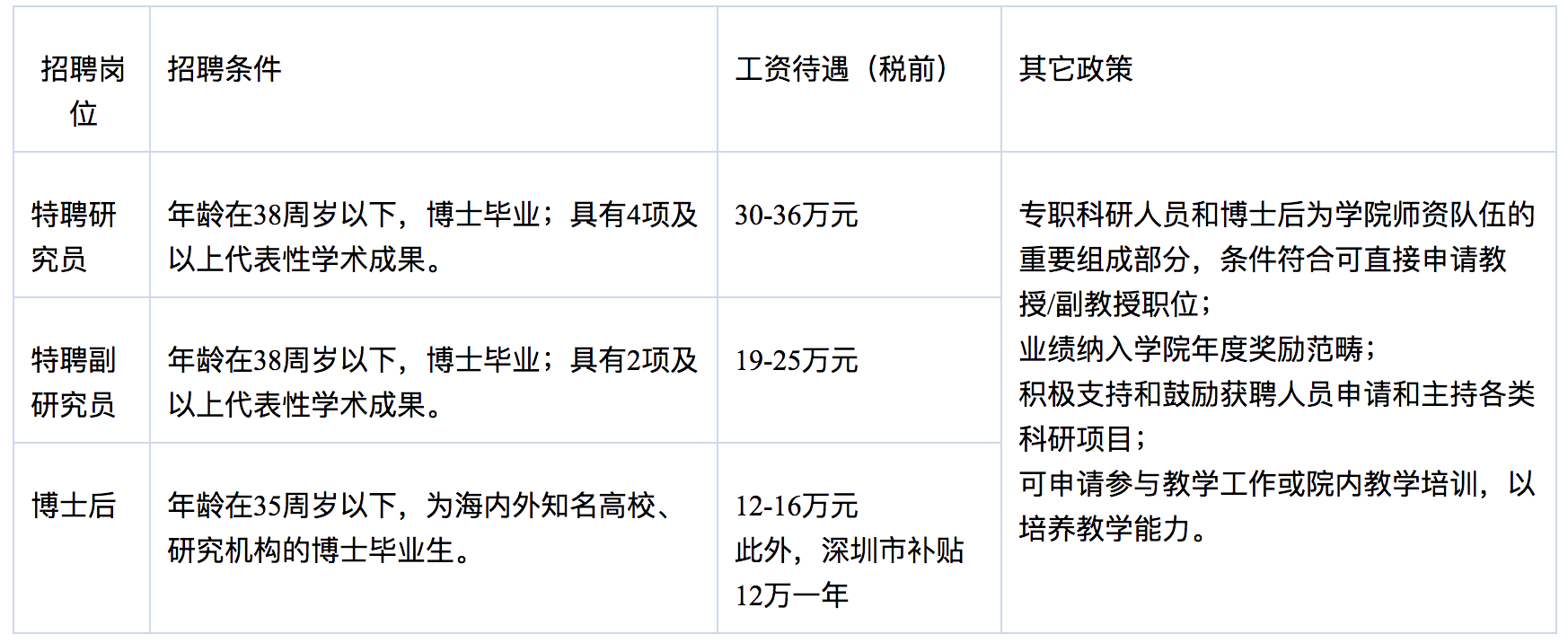
备注1:代表性学术成果主要为Thomson Reuters JCR-SCI 一区期刊论文,或中科院JCR-SCI 分区表二区期刊以上的论文。如主持国家级科研项目,或取得特别突出的学术成果(以第一作者或通讯作者在国际一流学术刊物或本学科公认最好的学术刊物上发表论文),代表性学术成果项数可放宽。
备注2: 专职科研人员无明确教学任务可申请各类科研项目,优秀者在聘期结束后或聘期内可直接申报学校的教授/副教授位置。 学校为录用并入职者报销参加面试的差旅费。
中山大学博士后人才项目
“博士后国际交流计划”派出项目:国家资助派出人员第一年每人30万人民币,第二年的资助经费由国外(境外)接收机构或合作导师承担。
“博士后国际交流计划”引进项目:资助标准为30万元人民币/人/年,其中国家资助20万元人民币/人/年,引进单位资助10万元人民币/人/年。
博士后创新人才计划:资助每人每年30万元,两年60万元,其中20万元为博士后科学基金。
香江学者计划:港方支付30万港币,国家资助30万元人民币。
广东省“珠江人才计划”海外青年人才引进计划:资助标准为每人两年60万元人民币。出站后连续在粤工作三年以上的博士后,给予每人40万元一次性安家补助。
申请材料及联系方式
有意者请将详细简历(包括学历、研究方向、发表文章等)通过电子邮件发至:lixin4306ren@gmail.com,请注明申请的职位。
本招聘启事长期有效,欢迎对生物医学研究有热情的青年才俊加盟!